SLAC joins the global fight against COVID-19
The lab is responding to the coronavirus crisis by imaging disease-related biomolecules, developing standards for reliable coronavirus testing and enabling other essential research.
By Manuel Gnida
As part of the international response to the COVID-19 pandemic, the U.S. Department of Energy’s SLAC National Accelerator Laboratory has re-opened some of its facilities for essential research on the atomic structure of the virus and how it interacts with potential treatments and vaccines.

SLAC joins the global fight against COVID-19
SLAC scientists are also leading the development of global standards to ensure reliable testing for the coronavirus, and they are participating in DOE working groups that are considering a wide range of proposals for coronavirus research, including high-throughput drug screening and novel approaches for building ventilators.
“Working with our fellow national labs, Stanford University and other partners, we’re applying our unique expertise and facilities to make a difference in the global fight against this disease,” said SLAC Director Chi-Chang Kao.
Targeting the virus on an atomic level
At the forefront of SLAC’s coronavirus research are techniques that use beams of X-rays or electrons to decipher how life’s molecular machinery works on the atomic level. This can help answer key questions about SARS-CoV-2, the virus that causes COVID-19, such as how it recognizes and infects cells, how it replicates within them, and how it spreads through our bodies. This detailed knowledge can feed into the development of vaccines and treatments with drugs or antibodies.
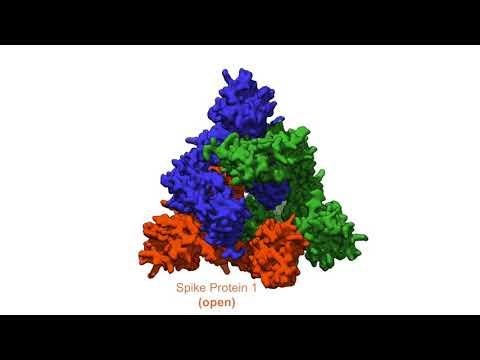
S-protein – ACE2
While the March 16 San Francisco Bay Area’s shelter-in-place order temporarily shut down all but essential operations at the lab, the Stanford Synchrotron Radiation Lightsource (SSRL) is operating a subset of its experimental stations to provide powerful X-rays for coronavirus-related studies.
One method they are using is X-ray crystallography, which examines biomolecules such as the spike proteins on the surface of SARS-CoV-2. These proteins dock onto receptor molecules on the surface of cells in the lungs, heart or intestines and spill the virus’s genetic material into the host cells where it replicates.
Researchers crystallize these biomolecules for analysis with SSRL’s X-rays, which produces detailed images of where each atom of the protein is located. These 3D maps can then feed into the design of targeted therapeutic interventions, for instance drugs that block spike proteins from binding to host cells.
Collaborating with researchers from other institutions, the SSRL team is working to determine the 3D structures of a number of biomolecules involved in the life cycle of SARS-CoV-2.

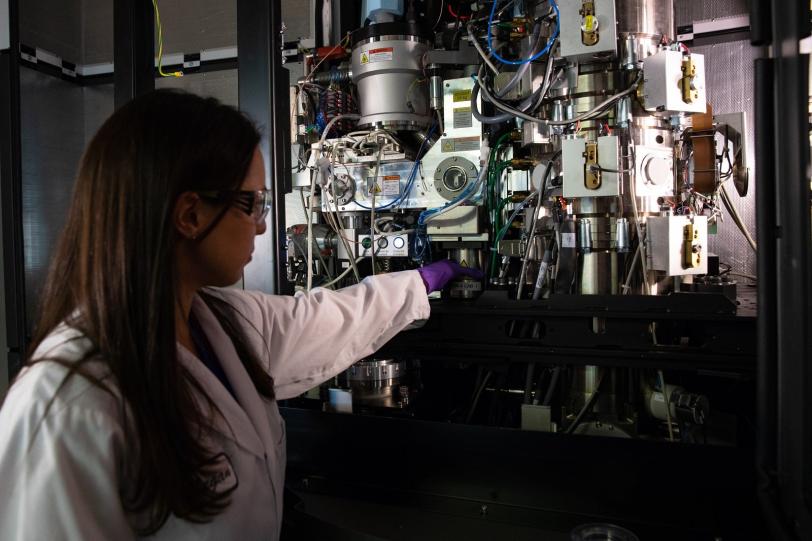
Robotic sample changer
“We’re able to do this research and, at the same time, follow strict social distancing protocols because our experimental stations are highly automated and require only minimal staff on site,” said Aina Cohen, who co-leads SSRL’s Structural Molecular Biology Division. “Researchers can send us frozen samples to be mounted by robots into the experimental stations. The remote researchers then use our software and video tools to control the robots, view their samples, and control all the details of their experiment from their home lab, or even from their homes.”
Another method used at SSRL is small-angle X-ray scattering, which looks at how X-rays bounce off biomolecules and their complexes in solution. Thomas Weiss, who oversees the research at this experimental station, is collaborating with Gerard Wong’s lab and other scientists at the University of California, Los Angeles, as well as researchers at the University of California, San Diego, to look at how pieces of viral proteins interact with the machinery of the human immune system. Insights into these interactions could help scientists understand why SARS-CoV-2 is so infectious and causes so much inflammation – a principal reason for why it is so deadly.

In parallel, researchers at two National Institutes of Health (NIH) centers – the Stanford-SLAC Cryo-EM Center and the National Center for Macromolecular Imaging – are pursuing answers to very similar questions with cryogenic electron microscopy (cryo-EM), which uses electron beams to study the novel coronavirus and how its molecular components interact with drugs or antibodies. One advantage of this technique is that it produces atomic-resolution images of biomolecules in their frozen, hydrated state, with no need for crystallization.
“One of the projects in full swing is a collaboration with Professor Rhiju Das at the Stanford Medical School that is aimed at designing a drug to keep viral RNA from carrying out its normal function,” said Wah Chiu, director of the NIH centers. This work involves developing synthetic RNA molecules that mimic stretches of the viral RNA, determining their 3D structures with cryo-EM and using that information to build molecules that can bind and deactivate the RNA.

In addition, Chiu and his colleagues have teamed up with several groups around the world that are preparing virus-like particles, viral proteins and RNA-protein complexes for cryo-EM analysis, as well as virus-infected cells for tomographic studies of what exactly the virus does once it enters host cells.
SLAC is also planning to bring experimental stations at its Linac Coherent Light Source (LCLS) back on line for COVID-19-related research. One of the brightest X-ray sources on Earth, this X-ray laser will complement other techniques – looking, for example, at hard-to-crystallize viral particles at room temperature to see how the virus attacks host cells and interacts with potential drugs under conditions that are close to those in the human body.
Toward reliable coronavirus testing
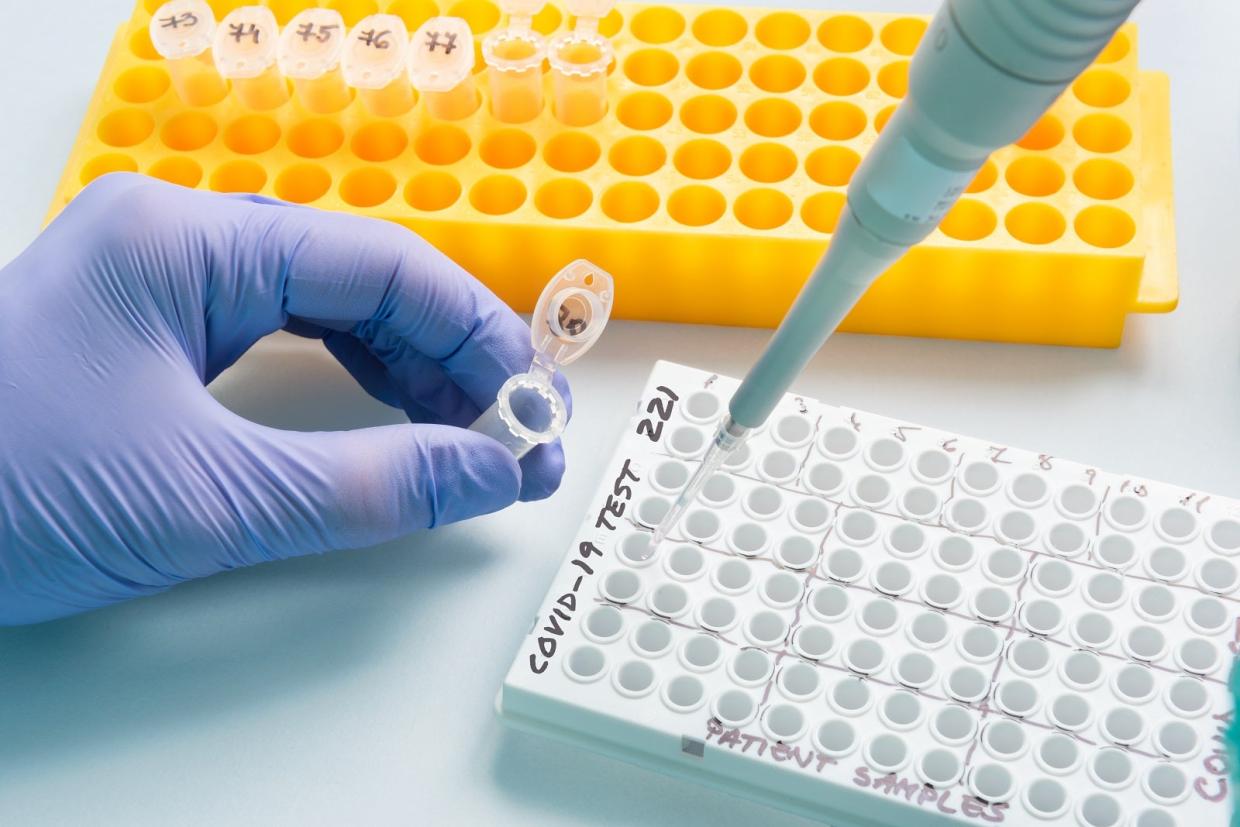
Another important area is making sure that tests for coronavirus are reliable, no matter when or where they take place. Defining standards that guarantee just that is the focus of an initiative launched by SLAC’s Marc Salit, director of the Joint Initiative for Metrology in Biology (JIMB).
“My team has been developing a Coronavirus Standards Working Group as the authoritative body to catalog existing standards and studies that will give us a global COVID-19 testing enterprise we can trust, and also fill in any gaps we find,” Salit said. “The idea is to create something enduring, institutionalized and poised for the next time so we can ensure we have the tools available to underpin the development of reliable testing.”
Although the working group is only a few weeks old, it has already become an active research community, bringing together leaders from government, academia, professional societies, international standards bodies, technology developers and the makers of standards and controls. The group is scoping standards for virus and antibody testing and exploring the creation of an open repository of clinical samples to support more test development and research.
A community-wide response
SLAC is also part of DOE’s COVID-19 working group, which brings together all 17 national labs in a coordinated response to explore the broad range of advanced scientific tools, deep technological knowledge and unique expertise the labs can bring to the table. This system-wide effort is developing ideas and approaches in a number of areas, including bioimaging, therapeutics, computing, materials and supply chains.
“SLAC is participating vigorously in all of these areas, leveraging our expertise in fabrication, electronics, prototyping, metrology and more,” said Steve Eglash, director of SLAC’s Applied Energy Division.
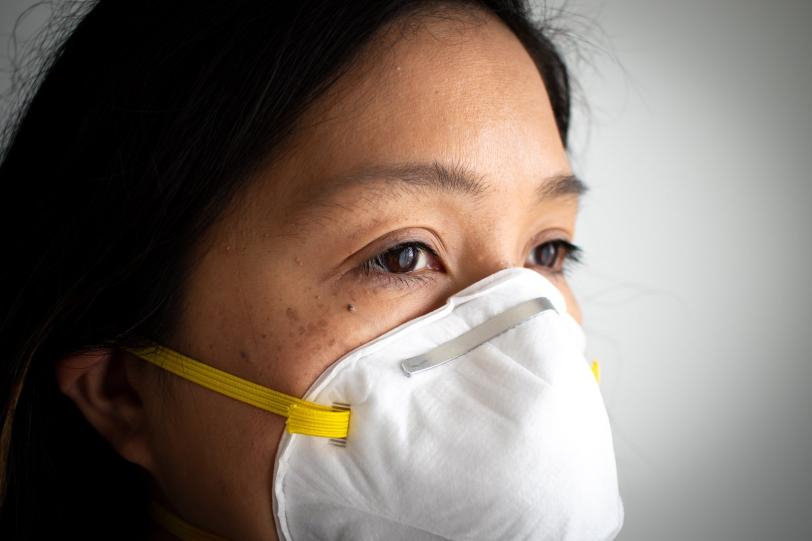

SLAC and Stanford faculty member Yi Cui, for example, has teamed up with Stanford colleague Steven Chu and Silicon Valley start-up 4C Air for a study that uses heat to sterilize N95 face masks that protect health care workers from COVID-19. The masks are in short supply, and disinfection could allow front-line medical workers to safely reuse them.
Other areas where SLAC’s expertise and tools could be used are the 3D printing of masks, the design of new, simplified types of ventilators, and the long-distance control of these devices, which would allow adjusting them without coming close to infectious patients. Members of SLAC’s machine learning initiative are also looking into how their computational methods can help with the analysis of complex cryo-EM and tomographic data.
“We’re seeing a lot of enthusiasm from many people all around the lab,” Eglash said. “Our community is really coming together and has already produced a long list of promising ideas to explore. The weeks and months ahead will tell us which of these projects will move ahead.”
Part of this work is funded by the DOE Office of Science. SSRL and LCLS are DOE Office of Science user facilities. Experimental stations for X-ray crystallography and X-ray scattering are part of the SSRL Structural Molecular Biology Program, which is supported by the DOE Office of Biological and Environmental Research and by the NIH, National Institute of General Medical Sciences. The Cryo-EM centers are funded by the NIH Common Fund Transformative High Resolution Cryo-Electron Microscopy Program and National Institute of General Medical Sciences Biomedical Technology Research Resource Program. JIMB is hosted by SLAC in collaboration with the National Institute of Standards and Technology (NIST) and Stanford.
For questions or comments, contact the SLAC Office of Communications at communications@slac.stanford.edu.
Editor's note: The studies described in this feature do not use infectious materials, such as live viruses.