Scientists make first high-res movies of proteins forming crystals in a living cell
A close-up look at how microbes build their crystalline shells has implications for understanding how cell structures form, preventing disease and developing nanotechnology.
By Glennda Chui
Scientists have made the first observations of proteins assembling themselves into crystals, one molecule at a time, in a living cell. The method they used to watch this happen – an extremely high-res form of molecular moviemaking – could shed light on other important biological processes and help develop nanoscale technologies inspired by nature.
Led by researchers at Stanford University and the Department of Energy’s SLAC National Accelerator Laboratory, the study was published in Nature Communications today.
“I’ve been super-excited to watch and track the movements of single molecules as they form this fascinating crystalline shell on the surface of a microbe,” said Stanford professor and study co-author W.E. Moerner, who shared the 2014 Nobel Prize in chemistry for stunning advances in pushing the boundaries of what optical microscopes can see. “We can look on a very fine scale and see the molecules arranging themselves in the shell. It’s the first time we’ve been able to do this.”
The study focused on a bacterium called Caulobacter crescentus that lives in lakes and streams. It’s one of many microbes that sport a very thin crystalline shell, known as a surface layer or S-layer, made of identical protein building blocks.
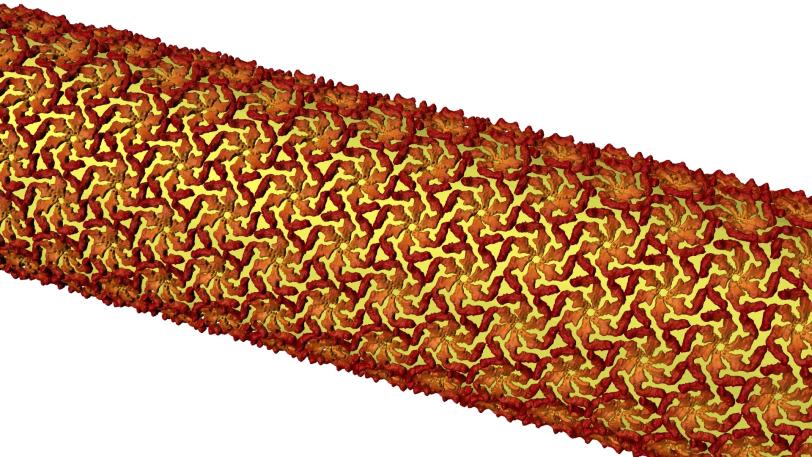

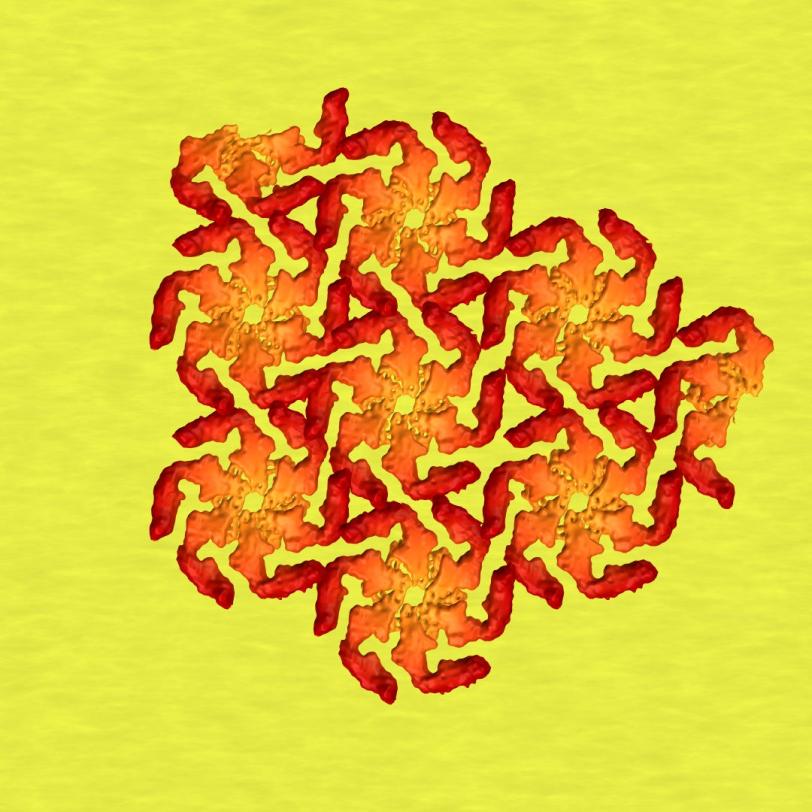
Scientists have been trying to figure out what roles these brittle shells play in the lives of their owners and how they come together to smoothly cover a microbe’s curvy surfaces. The research is driven not just by a desire to understand how nature works, but also by the possibility of applying that knowledge to create new types of nanotechnology – for instance, by using the protein shells as scaffolds for building “engineered living materials.” The shells also offer a potential target for drugs aimed at disarming infectious bacteria.
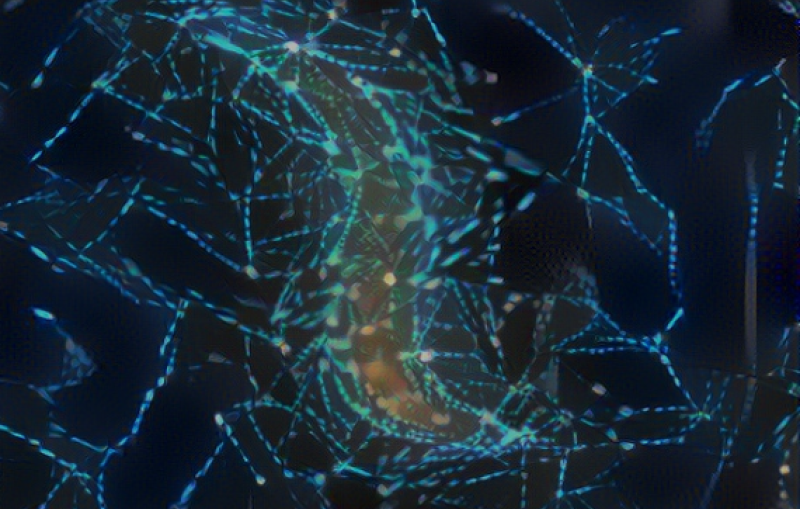
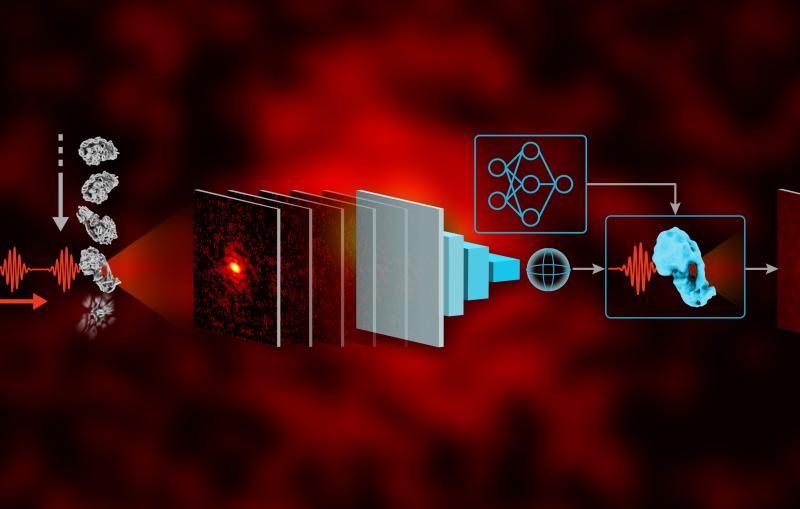
In this study, the research team used two established techniques that transcend the previous resolution limitations of optical microscopy – super-resolution fluorescence microscopy and single-molecule tracking – to watch individual building blocks move around the surfaces of living bacteria and assemble themselves into a shell. The resulting images and movies revealed how protein building blocks crystallize to form the bacterium’s S-layer coat.
“It’s like watching a pile of bricks self-assemble into a two-story house,” said Jonathan Herrmann, a PhD student at Stanford and SLAC who along with fellow Stanford PhD students Colin Comerci and Josh Yoon carried out the bulk of the work.
Protein crystallization
Following the glow
Protein crystals are widespread in nature: in shells that surround many bacteria and almost all of the ancient microbes called Archaea, in the outer shells of viruses and even in the human eye. The bacteria that cause anthrax and salmonella infections have these crystalline shells; so does Clostridium difficile, which causes serious infections of the colon and intestines. A lot of research has been aimed at disrupting these shells to head off infection.
The bacteria in this study don’t infect healthy people and are well-studied and understood, so they make good research subjects. Scientists know, among other things, that these bacteria can’t thrive without their shells, which are made from protein building blocks called RsaA. But shell assembly takes place at such a tiny scale that it had never been observed before.
To watch it happen, the researchers stripped microbes of their S-layers and supplied them with synthetic RsaA building blocks labeled with chemicals that fluoresce in bright colors when stimulated with a particular wavelength of light.
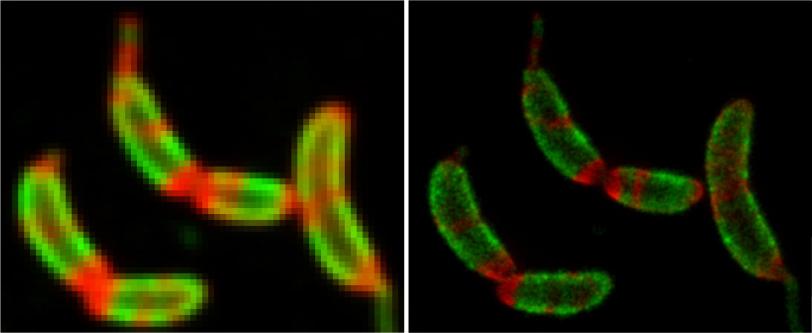
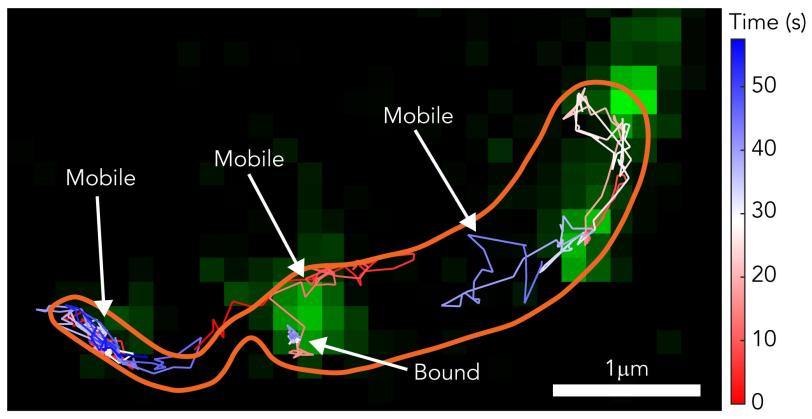
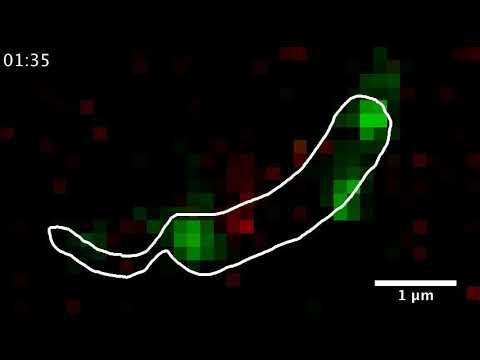
Then they tracked the glowing building blocks with single-molecule microscopy as they formed a shell that covered the microbe in a hexagonal, tile-like pattern in less than two hours. A technique called stimulated emission depletion (STED) microscopy allowed them to see structural details of the layer as small as 60 to 70 nanometers, or billionths of a meter, across – about one-thousandth the width of a human hair.
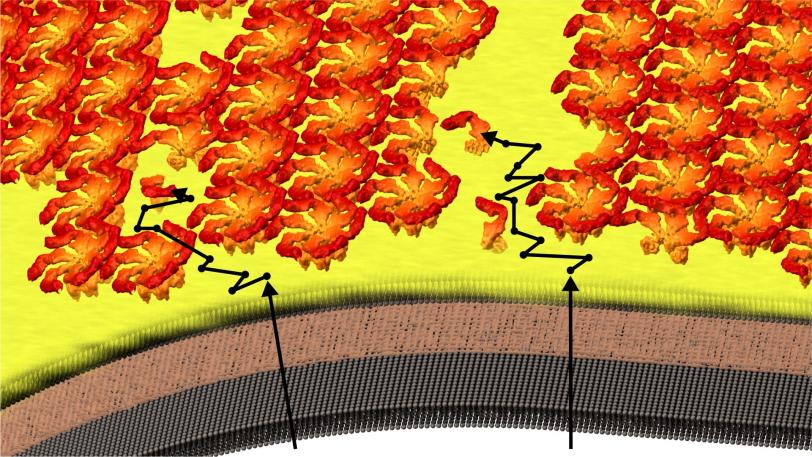
The team discovered that the shell-building didn’t happen the way they thought: The RsaA blocks were not guided into position and joined to the shell by enzymes, which promote most biological reactions. Instead they randomly moved around, found a patch of existing shell and joined it, like rock candy crystallizing around a string dipped in sugar water.
“The protein molecules are self-assembling building blocks, and they will spontaneously form themselves into crystals,” Herrmann said. “No enzyme is required.”
A new way of seeing
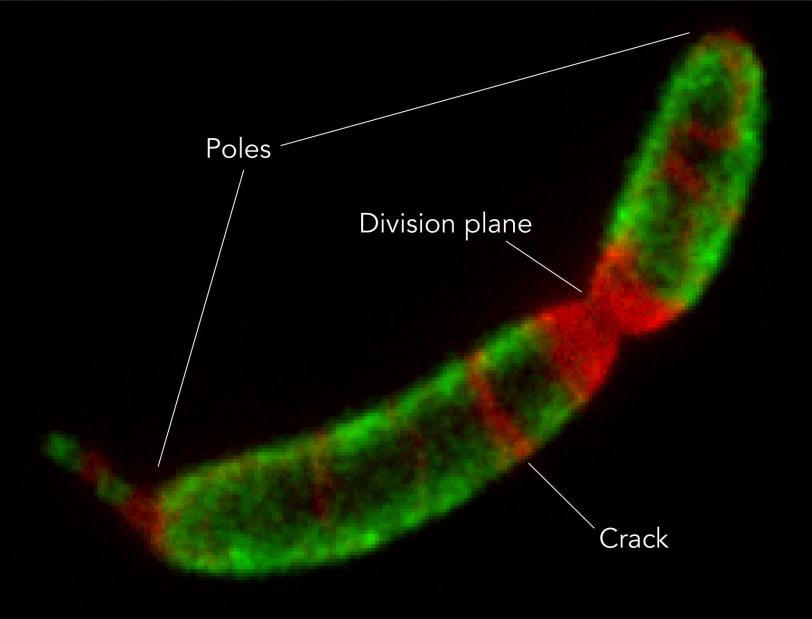
Since the flat crystalline shell can never perfectly fit the constantly changing 3-D shape of the microbe – “It’s not a huge leap to say that if you try to bend the sheet to fit the microbe, you have to break it,” Comerci said – there are always small defects and gaps in coverage, and those places, he said, are where they saw the shell grow.
“For the first time,” he said, “we were able to watch the S-layer proteins do things on their own.”

This new way of observing shell formation “is opening up a new way to understand and eventually manipulate surface layer structures, both in living organisms and in isolation,” said co-author Soichi Wakatsuki, a professor at SLAC and Stanford who leads the Biological Sciences Division at the lab’s Stanford Synchrotron Radiation Lightsource. “Now that we know how they assemble, we can modify their properties so they can do specific types of work, like forming new types of hybrid materials or attacking biomedical problems.”
The next step, researchers said, is to find out how the crystallization process starts using higher resolution X-ray and electron imaging available at SLAC: How do the very first bits of the shell crystallize without the equivalent of the rock candy string?
Optical microscopy for this study was carried out at the Moerner lab at Stanford. Researchers from the University of British Columbia and from Professor Lucy Shapiro’s laboratory at Stanford also contributed to this work, which was funded in part by the National Institute of General Medical Sciences and the Chan Zuckerberg Biohub. Work in Wakatsuki’s labs at SLAC and Stanford was partly funded by a Laboratory Directed Research and Development grant from SLAC and by the DOE Office of Biological and Environmental Research. The Stanford Synchrotron Radiation Lightsource is a DOE Office of Science user facility.
Citation: C. Comerci et al., Nature Communications, 21 June 2019 (10.1038/s41467-019-10650-x)
For questions or comments, contact the SLAC Office of Communications at communications@slac.stanford.edu.
About SLAC
SLAC National Accelerator Laboratory explores how the universe works at the biggest, smallest and fastest scales and invents powerful tools used by researchers around the globe. As world leaders in ultrafast science and bold explorers of the physics of the universe, we forge new ground in understanding our origins and building a healthier and more sustainable future. Our discovery and innovation help develop new materials and chemical processes and open unprecedented views of the cosmos and life’s most delicate machinery. Building on more than 60 years of visionary research, we help shape the future by advancing areas such as quantum technology, scientific computing and the development of next-generation accelerators.
SLAC is operated by Stanford University for the U.S. Department of Energy’s Office of Science. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time.