Researchers Take Snapshots of Potential 'Kill Switch' for Cancer
X-ray Study Shows Protein Switch for Programmed Cell Death in Motion
A study conducted in part at the Department of Energy’s SLAC National Accelerator Laboratory has revealed how a key human protein switches from a form that protects cells to a form that kills them – a property that scientists hope to exploit as a “kill switch” for cancer.
The protein, called cIAP1, shields cells from programmed cell death, or apoptosis – a naturally occurring crackdown on unhealthy cells and tissues. When a cell is in trouble, a signal activates cIAP1, which rapidly transforms into a state that allows apoptosis to take place.
“Cancer cells produce excess amounts of cIAP1 in an attempt to shut down apoptosis and evade death,” says senior staff scientist Thomas Weiss from SLAC’s Stanford Synchrotron Radiation Lightsource (SSRL), a DOE Office of Science User Facility, who participated in the study. “The search for drugs that would switch apoptosis back on to eradicate cancer is a very active research field.”
The researchers used X-rays from SSRL to watch in real time how cIAP1 transitions from one state to another. The results are an important step towards becoming able to control the protein’s switching properties.
“Our study closely ties cIAP1’s motions to its role as a switch,” says Allyn Schoeffler, a senior research associate at Genentech Inc. in South San Francisco and lead author of the study, published Nov. 10 in Nature Structural & Molecular Biology. “We now know why cIAP1 can act as a strictly controlled fail-safe for apoptosis and, at the same time, remain flexible enough to undergo rapid structural transitions.”
Incomplete Static Model
Earlier studies had given researchers a fairly good idea of cIAP1’s structure and general mechanism.
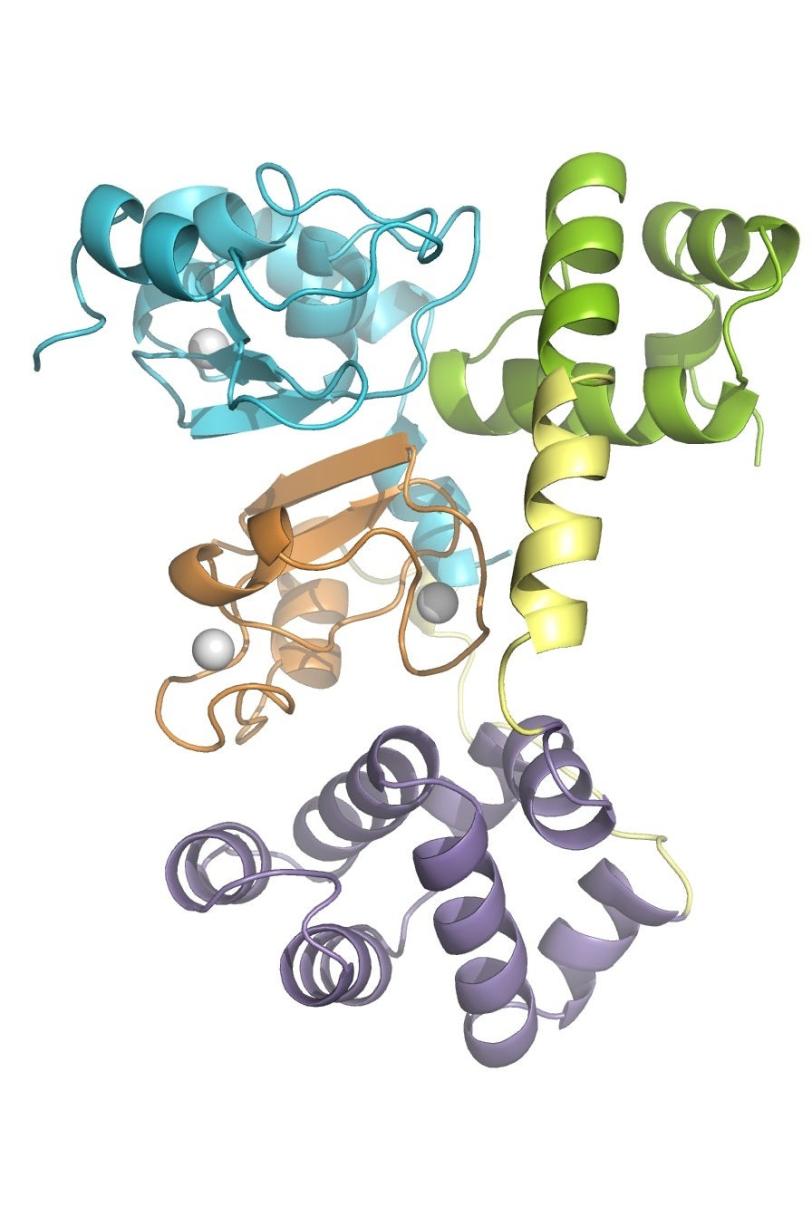
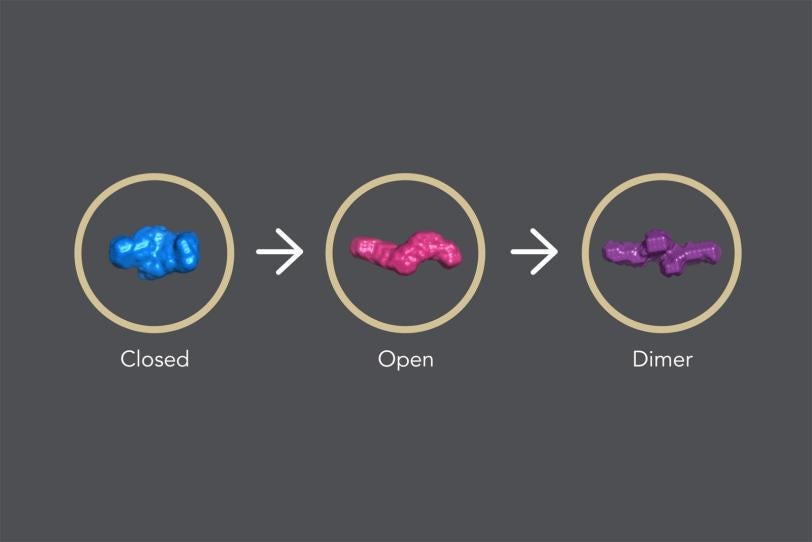
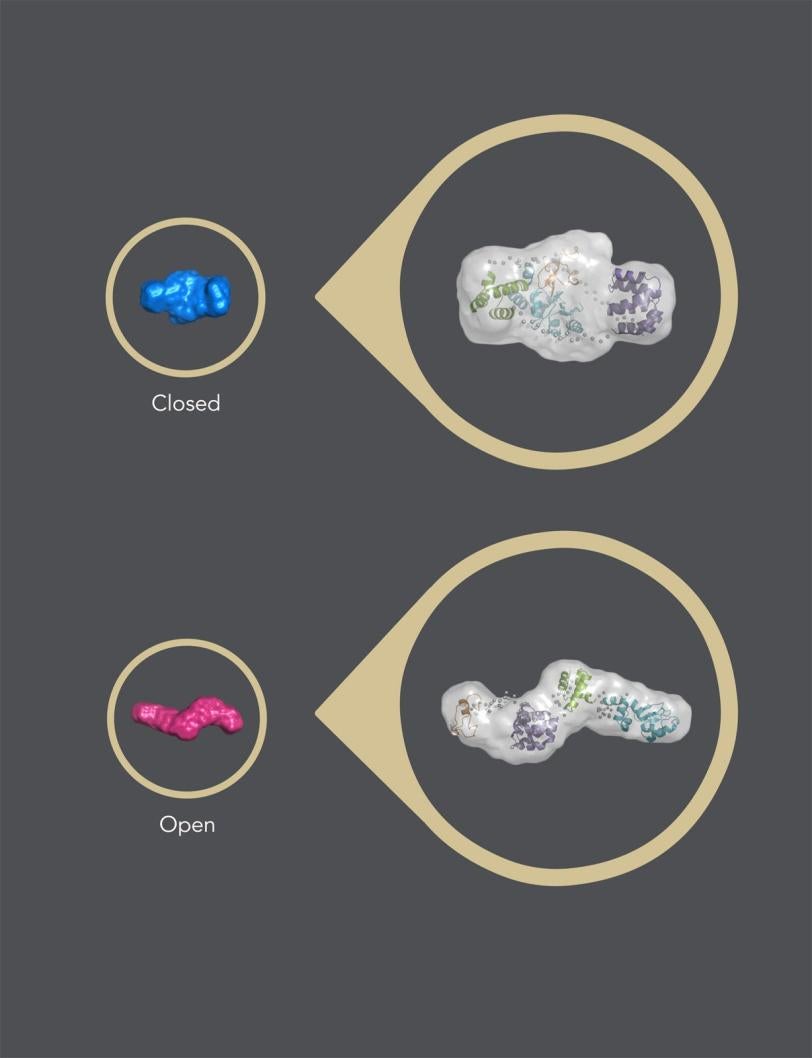
In its “closed” state, which blocks apoptosis, the protein’s four parts, or domains, are tightly bound together in a rather rigid, compact structure.
When a signal molecule binds to a specific site in cIAP1, the protein changes into its “open” state, in which the domains arrange in a more flexible, linear way. When two identical copies of this open structure partner up in what is known as a dimer, the assembly eventually self-destructs, removing the brake that blocks apoptosis and allowing cellular clean-up to carry on.
“This model of cIAP1 action has largely been derived from static images of the protein,” Schoeffler says. “However, static pictures do not tell us the whole story.”
Bringing Motion into the Equation
To find out more, the research team first used a technique known as nuclear magnetic resonance spectroscopy, or NMR, to analyze how the protein domains move in the closed state, and followed up with studies at SSRL, where they observed how X-rays scatter off the transforming sample.
“The results showed that cIAP1 switches from ‘closed’ to ‘open’ extremely fast, within only 300 milliseconds, which we were able to determine using a technique called time-resolved small-angle X-ray scattering,” says Weiss. “The following dimer formation is even faster than that.”
In addition, the scientists observed that the protein “breathes” rapidly in its closed form, with interfaces between domains opening and closing quickly.
“The only region that is relatively rigid is the interaction site for the apoptosis signal,” says Schoeffler. “This well-defined site in the closed state allows nature to control cIAP1 very tightly. It is the critical latch that keeps the switch closed and makes sure that it does not open accidentally.”
The rest of the protein, in contrast, is very flexible and allows cIAP1 to open instantaneously, like a spring-loaded trigger mechanism, when the proper signal is received. Once the trigger has been pulled, cell death becomes inevitable.
Ties to Cancer Research
The new insights could potentially benefit recent developments in cancer research. In fact, several studies are underway to explore the use of synthetic compounds that mimic nature’s signal molecules.
“Natural and synthetic molecules are thought to interact with this protein the same way,” says Schoeffler. “Therefore, the mechanisms revealed by our study are likely to hold true in medically relevant molecules as well.”
Research funding for the SSRL Structural Molecular Biology Program was provided by the DOE Office of Biological and Environmental Research and the National Institutes of Health, National Institute of General Medical Sciences.
Citation: A. Phillips, A. Schoeffler, et al., Nature Structural & Molecular Biology, 10 November 2014 (10.1038/nsmb.2916).
Contact
For questions or comments, contact the SLAC Office of Communications at communications@slac.stanford.edu.
SLAC is a multi-program laboratory exploring frontier questions in photon science, astrophysics, particle physics and accelerator research. Located in Menlo Park, Calif., SLAC is operated by Stanford University for the U.S. Department of Energy's Office of Science.
SLAC National Accelerator Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.