Fighting Ebola Virus Disease: 'Transformer' Protein Provides New Insights
Researchers have discovered that an Ebola virus protein can transform into three distinct structures with different functions. This rather uncommon property provides new clues for the development of potential drugs for deadly hemorrhagic fever.
By Manuel Gnida
A new study reveals that a protein of the Ebola virus can transform into three distinct shapes, each with a separate function that is critical to the virus’s survival. Each shape offers a potential target for developing drugs against Ebola virus disease, a hemorrhagic fever that kills up to 9 out of 10 infected patients in outbreaks such as the current one in West Africa.
At SLAC’s Stanford Synchrotron Radiation Lightsource (SSRL) and other X-ray facilities, a team led by Erica Ollmann Saphire of The Scripps Research Institute analyzed the structure of VP40, a protein best known for its role in creating and releasing new copies of the virus from infected cells.
“The interesting thing about VP40 is that it does more than that,” Saphire says. “We found that it is multifunctional, with several essential roles for the virus.” The team reported its results in Cell.
One Protein, Three Structures
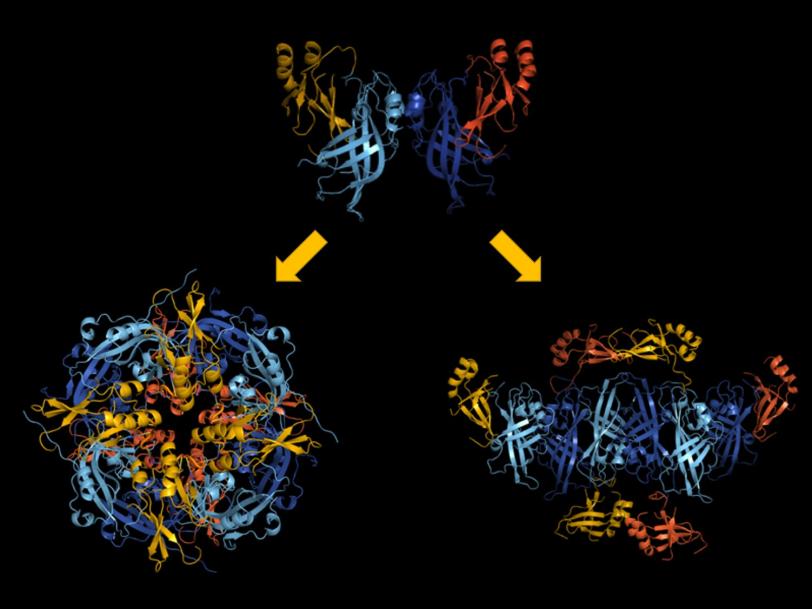
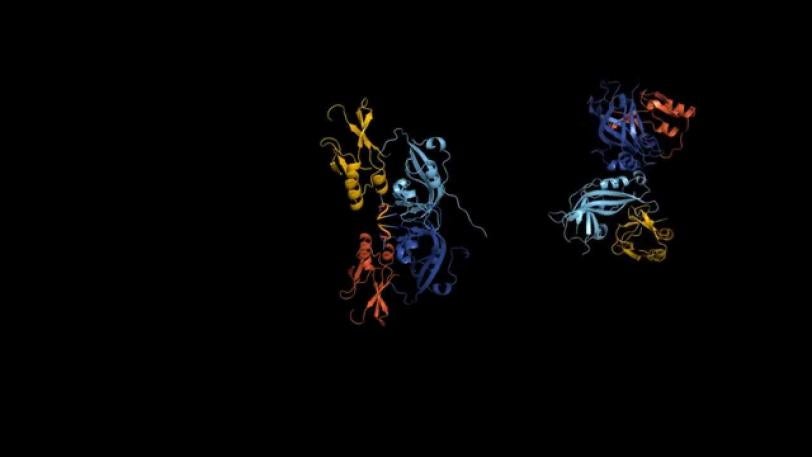
The team discovered that the protein can alter its shape, causing multiple copies of the protein to join up and create three very different assemblies: a butterfly shape composed of two, a ring formed by eight, and a linear structure built from six VP40 molecules. Prior to the study, only the protein ring was known.
But what unique functions do the individual structures have? The researchers took the study to the next level by combining their X-ray data with additional biological experiments. “This approach allowed us to track not only where the different structures are located, but also what they do inside the cell,” Saphire says.
It turns out that the function of each structure is linked to a specific stage of the virus life cycle.
While moving around inside infected cells, VP40 assumes the butterfly shape.
In the early stages of an infection, the VP40 molecules change their structure and assemble into a ring near the cell nucleus, regulating how the virus’s genetic information is copied.
In the later stages, VP40 travels to the cell’s outer layer, or membrane, and transforms into its linear structure, which plays a crucial role in the creation of new copies of the virus.
The transformational changes of VP40 update the nearly 60-year-old “central dogma of biology,” which implies that a given gene typically makes a single protein with a single 3-D shape. “Our findings open the central dogma wide up,” says Saphire, who suggests that structural rearrangements as seen in VP40 may be more common than previously thought.
From an evolutionary perspective, structural diversity has developed out of necessity. Unlike humans, who possess some 20,000 protein-encoding genes, the Ebola virus must get by with a drastically smaller number.
“The Ebola virus has only seven genes. However, its proteins must serve many more functions than that,” explains Scripps researcher Zachary Bornholdt, the study’s first author. “Protein transformability allows the virus to make the most out of very little.”
Potential Drug Targets
All three functions of VP40 – traveling inside infected cells, regulating genetic information and creating new viruses – are essential to the Ebola virus, and disrupting any of the corresponding structures or their transformations would severely affect it. Therefore, VP40’s triple role provides researchers with important clues for the development of potential antiviral drugs.
“The more we are able to define VP40’s structures and functions, the more we can expand what we can do with this information,” Bornholdt says. “Our data suggest, for instance, that it might be more effective to target the ring than the other structures because only a small fraction of all VP40 molecules form the ring in the course of the viral life cycle.”
Although a cure for Ebola virus disease is still remote, the new study already has practical applications: The same VP40 proteins produced for this study are being used in test strips to identify the disease in patients affected by the current outbreak in West Africa.
The research team included scientists from The Scripps Research Institute and the University of Wisconsin-Madison in the U.S., as well as from the University of Tokyo and the Exploratory Research for Advanced Technology Program in Japan. Part of the research was performed at SSRL's microbeam facility for crystallography (Beam Line 12-2). The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences.
Citation: Bornholdt et al., Cell 154, 763–774 (2013), DOI: 10.1016/j.cell.2013.07.015.
SLAC is a multi-program laboratory exploring frontier questions in photon science, astrophysics, particle physics and accelerator research. Located in Menlo Park, Calif., SLAC is operated by Stanford University for the U.S. Department of Energy's Office of Science.
SLAC's Stanford Synchrotron Radiation Lightsource (SSRL) is a third-generation light source producing extremely bright X-rays for basic and applied science. A DOE national user facility, SSRL attracts and supports scientists from around the world who use its state-of-the-art capabilities to make discoveries that benefit society. For more information, visit ssrl.slac.stanford.edu.
DOE's Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.