Strange Discovery: Bacteria Built with Arsenic
In a study that could rewrite biology textbooks, scientists have found the first known living organism that incorporates arsenic into the working parts of its cells.
By Kelen Tuttle
Menlo Park, Calif. — In a study that could rewrite biology textbooks, scientists have found the first known living organism that incorporates arsenic into the working parts of its cells. What's more, the arsenic replaces phosphorus, an element long thought essential for life. The results, based on experiments at the Stanford Synchrotron Radiation Lightsource, were published online today in Science Express.
"It seems that this particular strain of bacteria has actually evolved in a way that it can use arsenic instead of phosphorus to grow and produce life," said SSRL Staff Scientist Sam Webb, who led the research at the Department of Energy's SLAC National Accelerator Laboratory. "Given that arsenic is usually toxic, this finding is particularly surprising."
Phosphorus forms part of the chemical backbone of DNA and RNA, the spiraling structures that carry genetic instructions for life. It is also a central component of ATP, which transports the chemical energy needed for metabolism within cells. Scientists have for decades thought that life could not survive without it.
But this was not the case for a strain of Halomonadaceae bacteria called GFAJ-1, found in an eastern California lake. Colonies of these bacteria flourished, as expected, when given a steady supply of phosphorus along with other necessities; yet when researchers replaced the phosphorus with arsenic, the colony continued to grow.
This suggested to Felisa Wolfe-Simon, a NASA research fellow and geobiologist in residence with the U.S. Geological Survey, that the bacteria were using the arsenic in place of phosphorus.
"We already knew that other microbes can 'breathe' arsenic, but it seemed these bacteria could be doing something new: building parts of themselves out of arsenic," said Wolfe-Simon, the paper's lead author. "To see if that was the case, we brought samples to SSRL. I came armed with the knowledge that the bacteria were doing something really weird, and I knew that SSRL beamline 2-3, in Sam's hands, could tell us more."
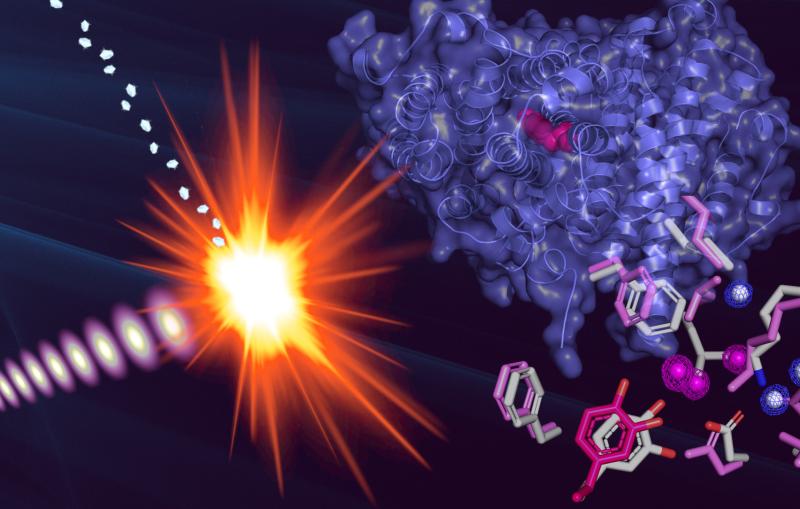
Wolfe-Simon's overall goal was to see if the arsenic was intimately associated with the bacterial cells or simply attached to the outside. The team swept SSRL's hair-thin X-ray beam across a sample of bacteria that had been bathed in high concentrations of arsenic. The interaction between the X-rays and the sample revealed where and how the arsenic wound up inside the bacterial cells.
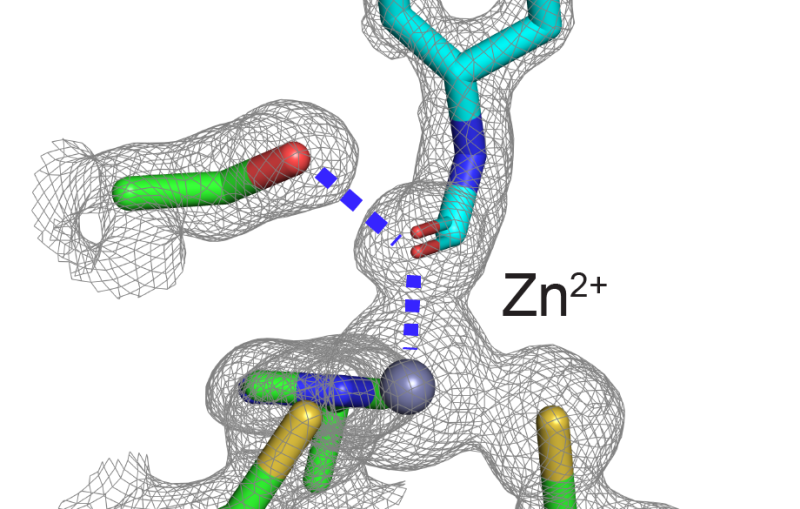
"We saw similarities in the distribution of arsenic and the distribution of iron and zinc," two metals that indicate where an organism's cellular material is located, Wolfe-Simon said. In contrast, the distribution of phosphorus didn't match the distribution of the other elements, suggesting that arsenic took the place of phosphorus in the bacteria's cellular material.
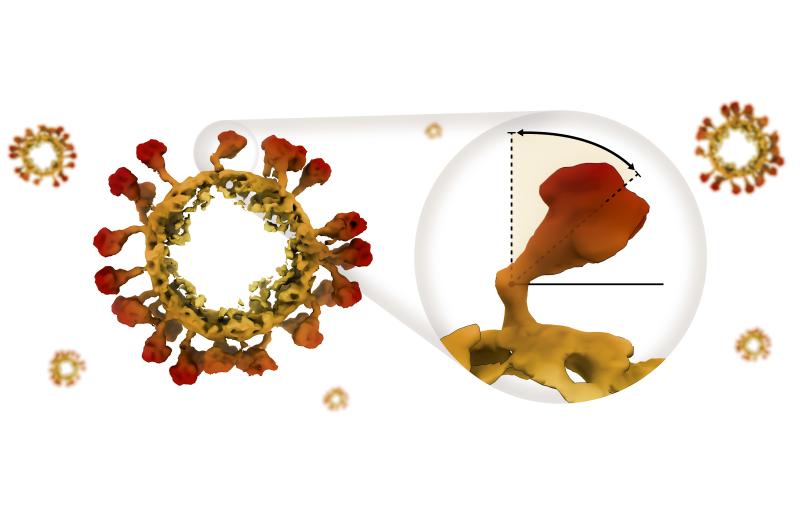
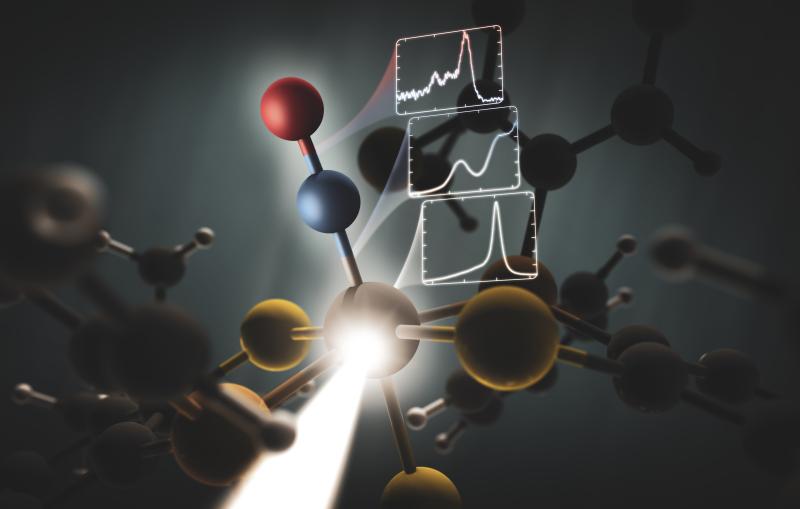
To confirm this suspicion, the team conducted another experiment with the SSRL beam, this time in a spectroscopic mode, which identifies the types and locations of specific atoms. The experiment revealed that the arsenic atoms' nearest neighbors were oxygen and, at slightly longer distances, carbon. This pattern and the specific distances between the three types of atoms are a near-perfect match with the way phosphorus atoms bind to other atoms in strands of conventional DNA.
Wolfe-Simon said that these experiments strongly suggest that the bacteria aren't just absorbing arsenic, but incorporating it into their own beings as "biological arsenic."
Further bolstering this view, the arsenic did not take the form that would be expected if, for example, an organism were trying to remove the toxin from its system, and it was not surrounded by the types of molecules that an organism might use to render the arsenic inert, Webb said. The fact that it was attached to carbon and oxygen, he said, is "what we would expect if it were actually being used to create DNA, RNA or proteins." This would make the bacterium strain GFAJ-1 the first organism known to use arsenic in place of phosphorus for growth.
"In theory, this knowledge will rewrite biology textbooks," Wolfe-Simon said. "Whenever you hear about 'diversity' in biology, it always means metabolic diversity—diversity in what organisms breathe or what they oxidize, how they make a living. It's assumed that however diverse organisms may be, they're all made of the same elements: carbon, hydrogen, nitrogen, oxygen, phosphorus and sulfur. This type of bacterium is not. And that suggests that there may be a whole new world of organisms to explore."
Next, the team plans to investigate specific ways these bacteria might use arsenic in proteins, fats and nucleic acids such as DNA.
"To do this, we'll need a bigger sample," said Webb. "Felisa and her team are working to grow that now, and we're looking forward to further studying the details of this amazing organism at SSRL."
This research was conducted by Felisa Wolfe-Simon (NASA Astrobiology Institute, U.S. Geological Survey), Jodi Switzer Blum (U.S. Geological Survey), Thomas R. Kulp (U.S. Geological Survey), Gwyneth W. Gordon (Arizona State University), Shelley E. Hoeft (U.S. Geological Survey), Jennifer Pett-Ridge (Lawrence Livermore National Laboratory), John F. Stolz (Duquesne University), Samuel M. Webb (SSRL and SLAC), Peter K. Weber (Lawrence Livermore National Laboratory), Paul C.W. Davies (NASA Astrobiology Institute, Arizona State University), Ariel D. Anbar (NASA Astrobiology Institute, Arizona State University), and Ronald S. Oremland (U.S. Geological Survey).
SLAC National Accelerator Laboratory is a multi-program laboratory exploring frontier questions in photon science, astrophysics, particle physics and accelerator research. Located in Menlo Park, California, SLAC is operated by Stanford University for the U.S. Department of Energy Office of Science. The Stanford Synchrotron Radiation Lightsource at SLAC is a DOE Office of Science national user facility which provides synchrotron radiation for research in chemistry, biology, physics and materials science to more than a thousand users each year. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Science and by the National Institutes of Health, National Center for Research Resources' Biomedical Technology Program.
Contact
For questions or comments, contact the SLAC Office of Communications at communications@slac.stanford.edu.